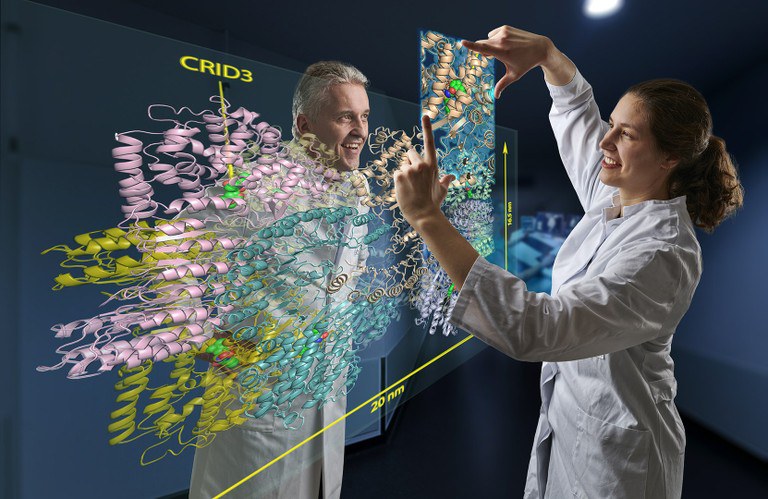
In their study, the scientists examined a protein molecule with the cryptic abbreviation NLRP3. This is a kind of danger sensor in the cell: it sounds the alarm when the cell comes under stress, for example due to a bacterial infection or toxins.
NLRP3 then causes holes to form in the cell membrane, which ultimately leads to the death of the cell. Before this happens, however, the sensor molecule stimulates the formation of inflammatory messenger substances that are released through the perforated membrane. They recruit further immune cells to the site of the incident and ensure that cells in the vicinity commit suicide - so that a bacterium or virus, for example, cannot spread any further.
"The result is a massive inflammatory reaction," explains Prof. Dr. Matthias Geyer from the Institute of Structural Biology at the University of Bonn, who led the study. "For defense against pathogens, this is also very useful. But if it is overdosed or already triggered by harmless triggers, chronic inflammatory diseases can be the result - such as type II diabetes, gout, Crohn's disease or even dementia diseases like Alzheimer's."
Targeting inflammation
Around the globe, researchers are therefore looking for ways to target inflammatory processes without undermining the entire mechanism of the immune response. Twenty years ago, the U.S. pharmaceutical company Pfizer published an interesting finding in this regard: certain active ingredients prevent the release of cytokines, the most important inflammatory messengers. How these CRIDs (Cytokine Release Inhibitory Drugs) do this, however, was unknown until now.
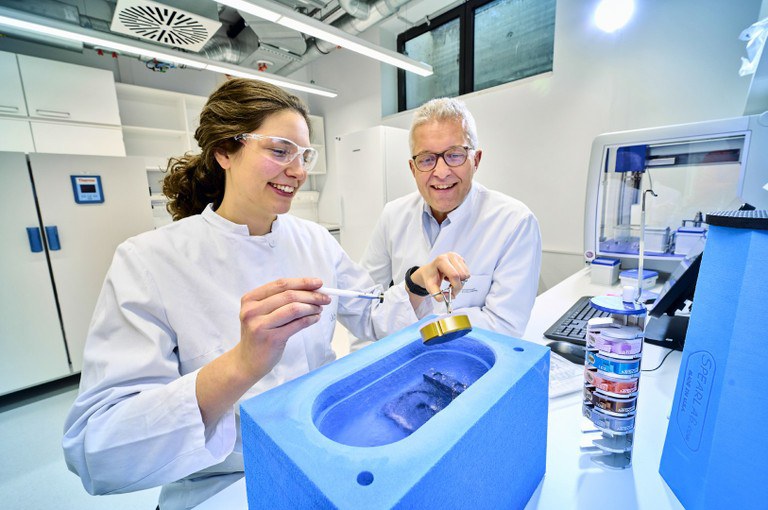
It has been known for several years that CRIDs somehow prevent cellular danger sensors from sounding the alarm. "We have now discovered the way in which they exert this effect," explains Geyer's colleague Inga Hochheiser. To do this, she isolated large amounts of NLRP3 from cells, purified them and mixed them with the inhibitor CRID3. She dropped tiny portions of this mixture onto a carrier and then froze them directly.
This method produces a thin ice film. It contains millions of NLRP3 molecules to which CRID3 is bound. These can be viewed with an electron microscope. Because the molecules fall differently as they drop, different sides of them can be seen under the microscope. "These views can be combined to form a three-dimensional image," Hochheiser explains.
The cryo-EM images allow a detailed look at the structure of the hazard sensor inactivated by CRID3. They show that NLRP3 in its inactive form assembles into a mega-molecule. It consists of ten NLRP3 units that together form a kind of cage. "But the most exciting result of our work is that we were able to identify the CRID3 binding sites," Geyer is pleased to report. "All other research groups have cut their teeth on this so far."
Inhibitor prevents unfolding of giant molecule
The binding sites (structural biologists also speak of "pockets") are thus located inside the cage. Each of the ten units has one of these pockets. If it is occupied by CRID3, the inhibitor blocks a folding mechanism that is necessary for the activation of NLRP3. Similar to a blossoming rose, which can only be visited by a bee in this state, certain parts of the NLRP3 protein reach the surface of the cage when it is folded over and thus become accessible.
NLRP3 is a representative of a whole family of similar proteins. Each of them presumably performs its own specific task in different inflammatory processes. "Based on our studies, we believe that the pockets of all these NLRPs differ," Geyer says. "Therefore, a separate inhibitor can probably be found for each one." Researchers are thus provided with a whole arsenal of potential new weapons against various, inflammatory diseases.
For example, the current work makes it possible to search specifically for more effective alternatives to CRID3 that also have fewer side effects. But this is just the beginning, says Geyer, who is also a member of the ImmunoSensation2 cluster of excellence at the University of Bonn. "I'm convinced that our study will open up a fruitful new field of research that will keep scientists busy for decades to come."
Participating institutions and funding:
The study was supported by the German Research Foundation (DFG) and by EU funds under the iNEXT-Discovery and Instruct-ERIC initiatives. The cryo-EM images for structural elucidation were acquired at EMBL in Heidelberg.
Publication: Inga V. Hochheiser, Michael Pilsl, Gregor Hagelueken, Jonas Moecking, Michael Marleaux, Rebecca Brinkschulte, Eicke Latz, Christoph Engel & Matthias Geyer: Structure of the NLRP3 decamer bound to the cytokine release inhibitor CRID3; Nature; DOI: 10.1038/s41586-022-04467-w;
URL: https://www.nature.com/articles/s41586-022-04467-w
Contact:
Prof. Dr. Matthias Geyer
Institute for Structural Biology at the University of Bonn
Tel. +49-228/287-51400
E-Mail: matthias.geyer@uni-bonn.de